MONO-IRON HYDROGENASE
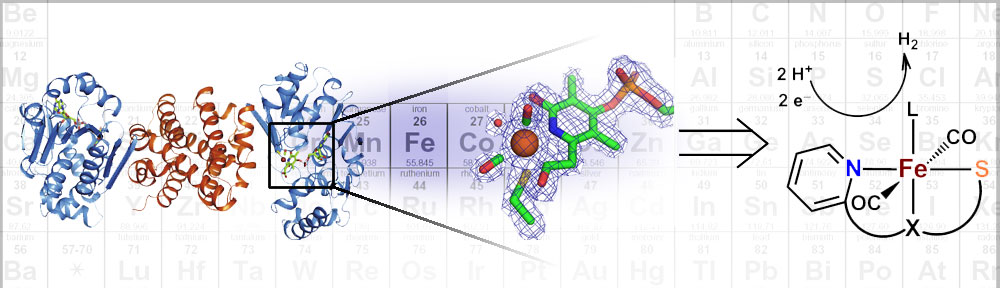
above image credit: Ermler et al, FEBS Letts, 2009, 583, 585
Background. The generation of dihydrogen (H2) as a chemical fuel is desirable as a sustainable and carbon-free fuel source. In fact, H2 can be generated from simple precursors: H2O and electrons. Currently, metallic platinum (Pt) is the best and most widely utilized catalyst for H2 generation. However, the low earth abundance and corollary high cost of Pt prohibits widespread, delocalized use of such a process.
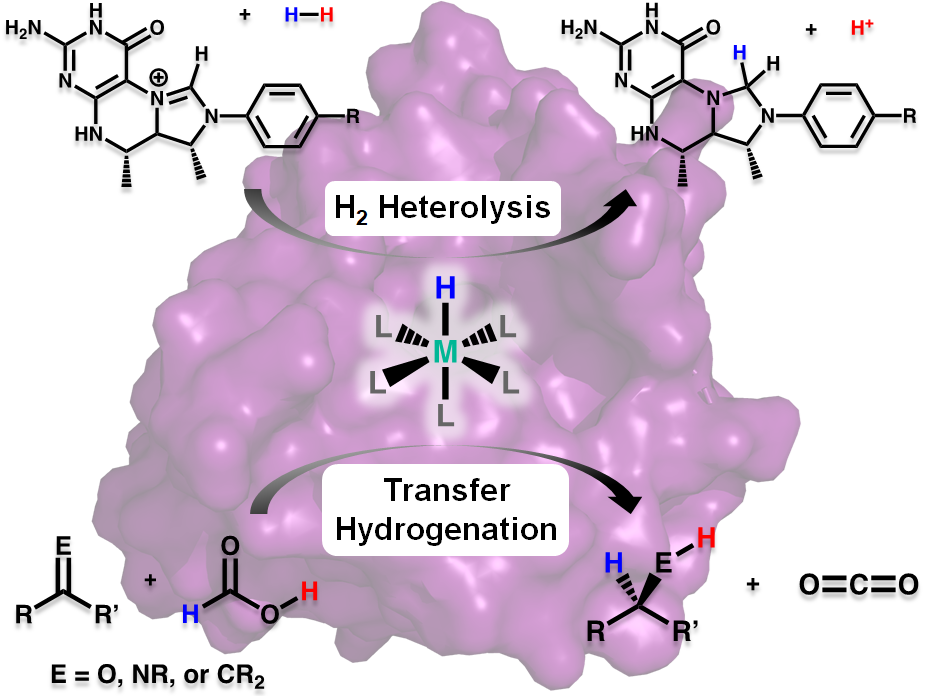
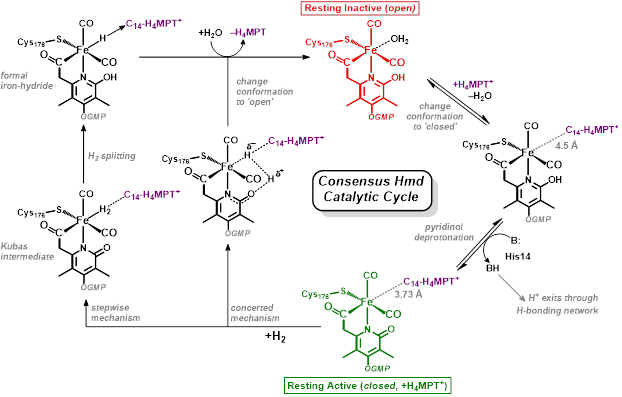
Research in the Rose group is inspired by one of nature’s H2 utilizing enzymes: Mono-Iron Hydrogenase, often abbreviated HMD-H2ase. The enzyme is also known as “cluster-free” hydrogenase due to its lack of an [Fe4S4] redox cluster (unlike the di-iron and nickel-iron hydrogenases). Of all the transition metals, iron is the cheapest and most earth abundant. Therefore development of an inexpensive, reliable, Fe-based catalyst could alter the landscape of H2 production.
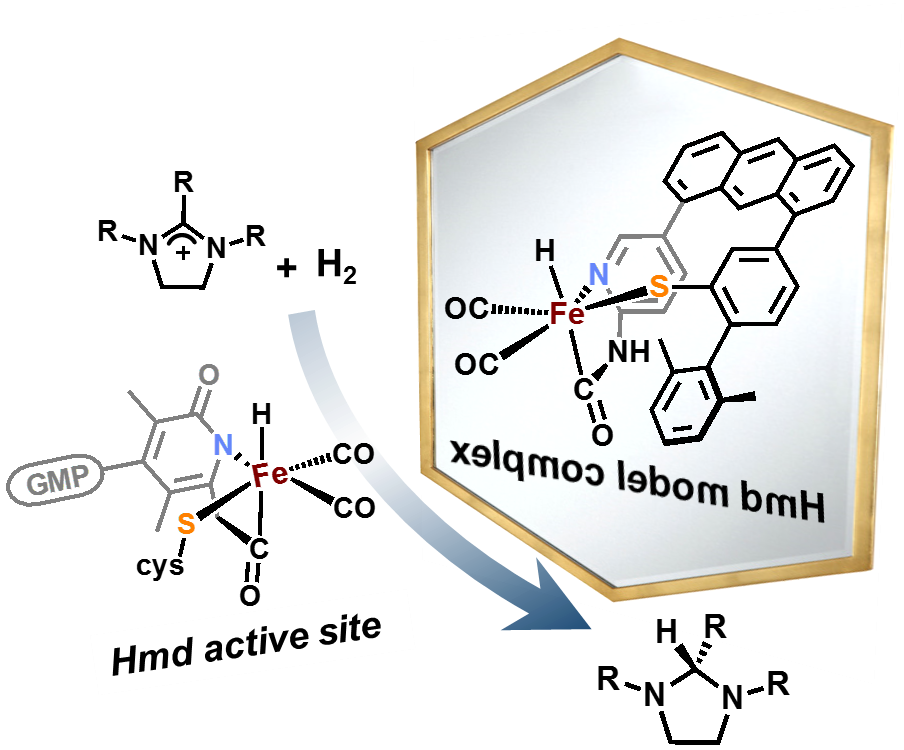
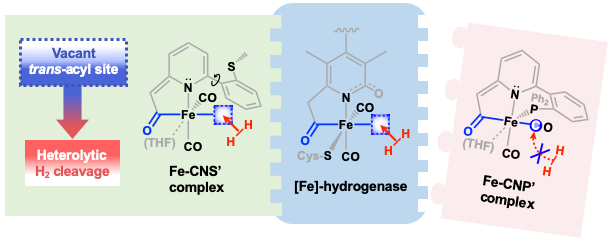
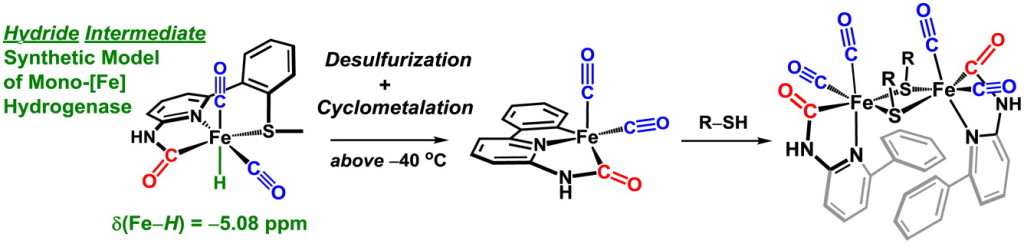
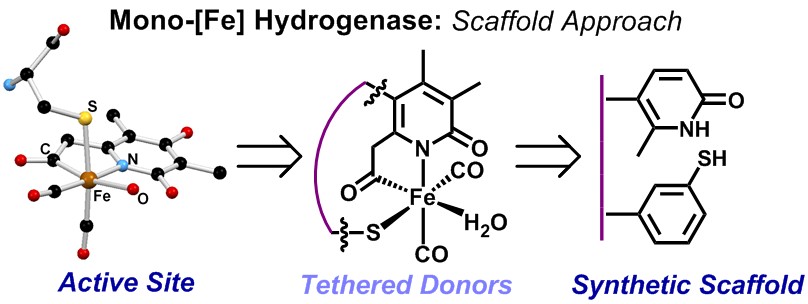
Our group uses the crystallographically characterized mono-Fe-H2ase active site as inspiration to prepare small molecule mimics of the enzyme.The enzyme active site presents a unique array of donors to the Fe center not yet found anywhere else in nature, including an N-bound pyridone cofactor and an unusual Fe-C(acyl) bond.
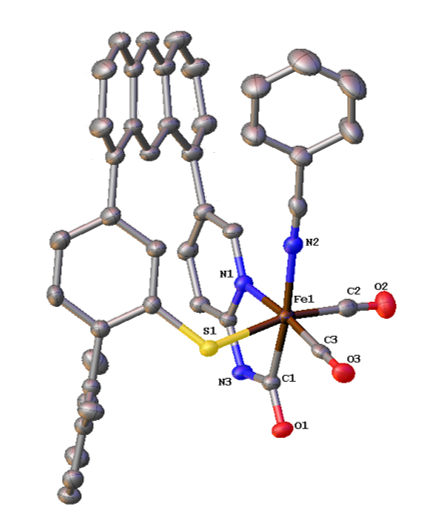
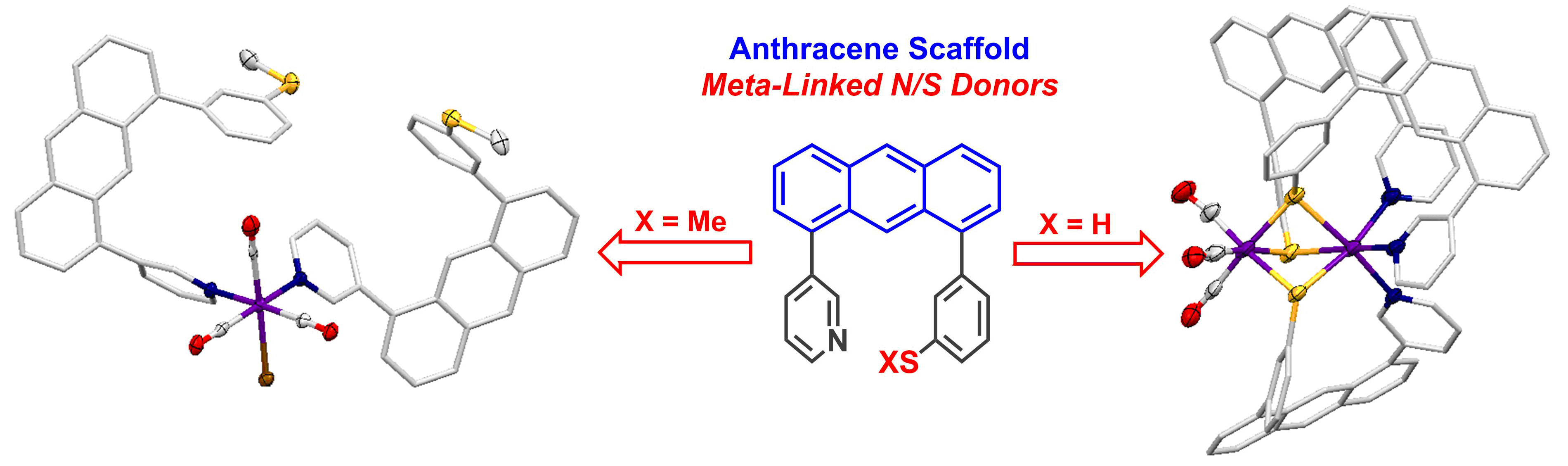
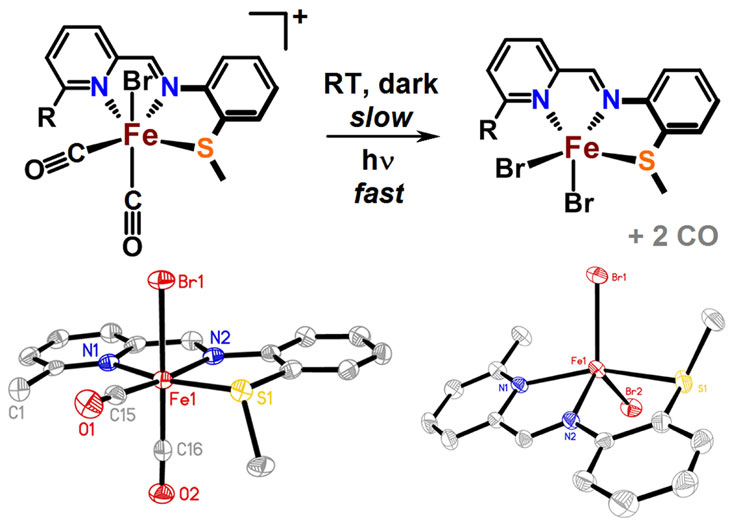
Scaffold-Based Ligand Design & In/Organic Synthesis. The rational design of scaffold-based ligands is a primary focus within the project. We use the resulting Fe complexes as structural and spectroscopic models to understand the properties and function of the Fe-H2ase active site. The wide variety of donor types and ligand architectures available through chemical synthesis is one particular advantage compared to the standard amino acids in a protein-based biochemical approach.
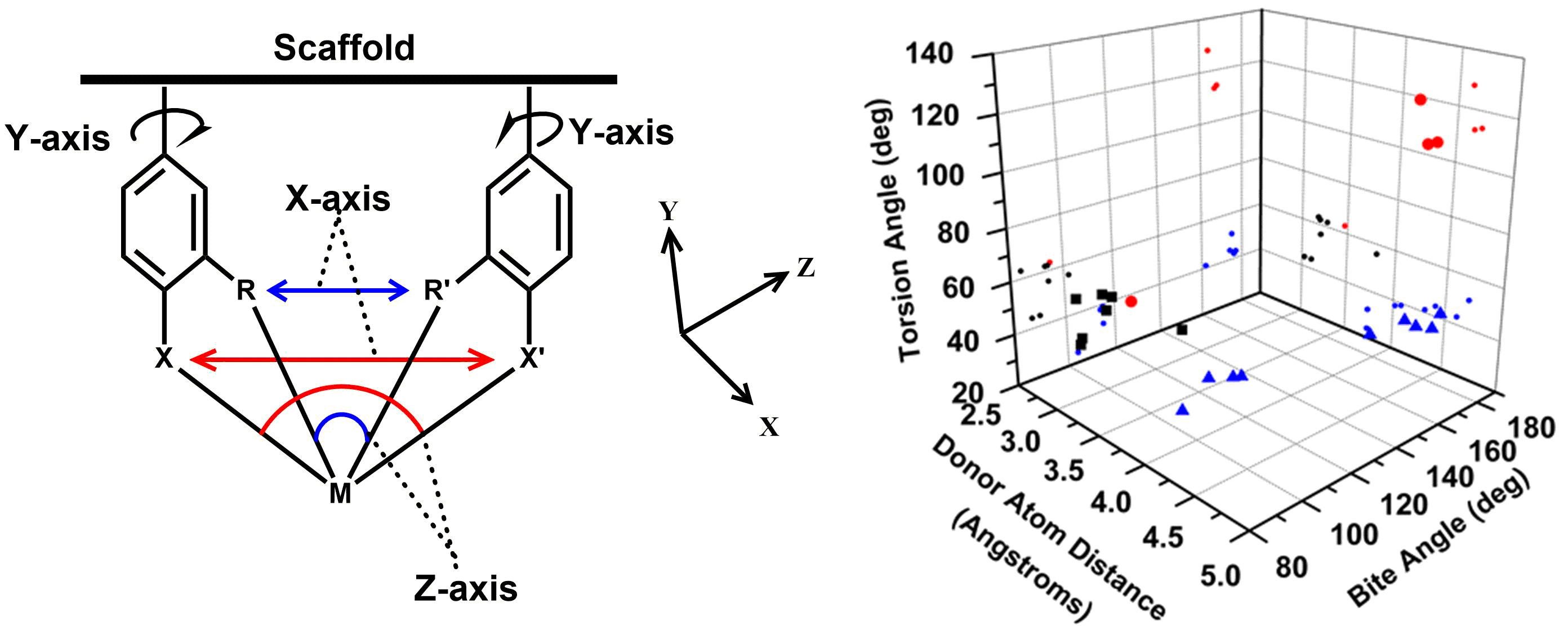
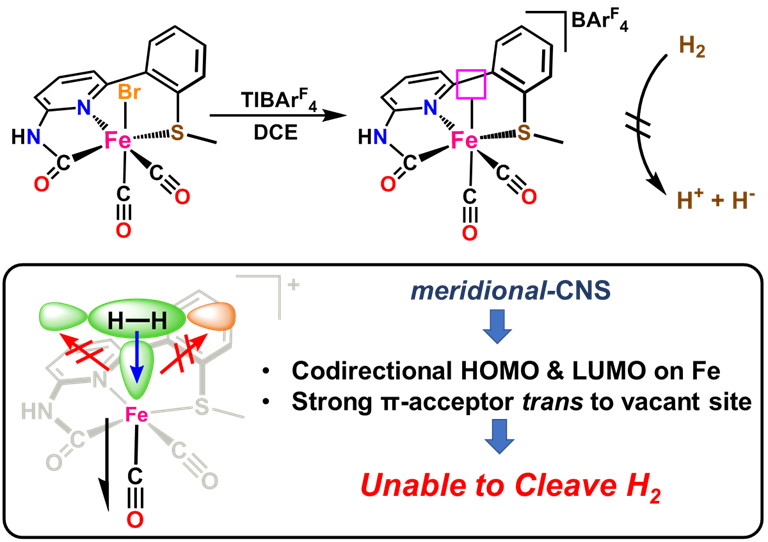
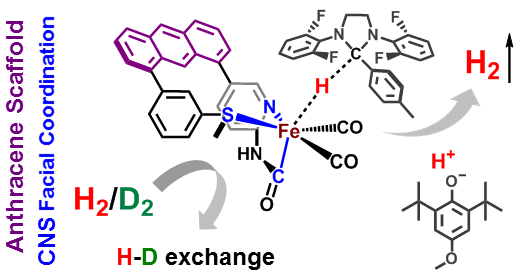
Dynamics & Flexibility. More recently, we have become interested in using our unique scaffold-based ligand design approach to probe the importance and effect of molecular dynamics and flexibility on reactivity at the metal center. In nature, proteins and metalloenzyme active sites are dynamic entities, and yet small molecule catalysts are historically rigid and non-dynamic. Much in the same way that the “second coordination sphere” of H-bonds and ionic contacts has been proven to be critical for understanding and modulating metal center reactivity, we believe that the “thematic third coordination sphere” is the extent of dynamic motion around the metal center that tunes its reactivity. The projects thus involve the synthesis of various flexible / dynamic scaffolds, preparation of Fe complexes thereof, and studying reaction rates and H2/D2 effects on kinetics to quantify the effect of dynamics on metal site reactivity.
NITROGENASE
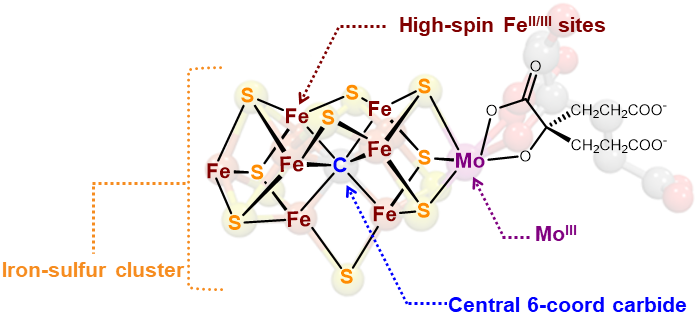
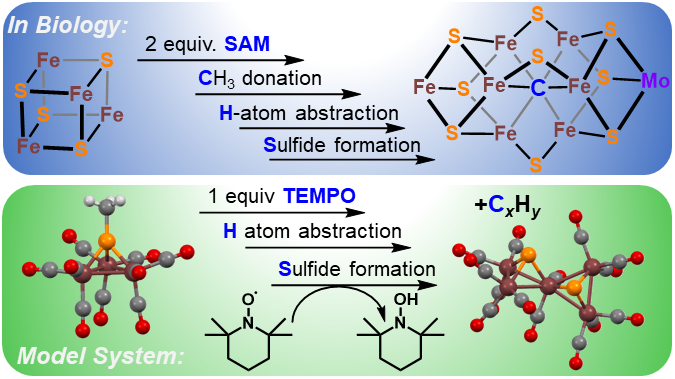
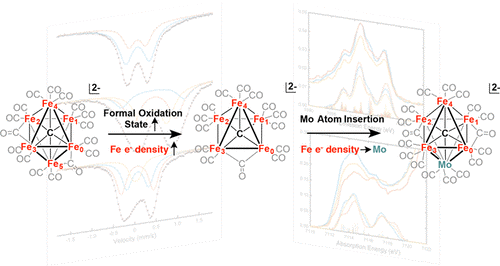
Background. Another synthetic modeling goal is the FeMoco active site of the enzyme nitrogenase. This fascinating enzyme contains a 6-coordinate, interstitial inorganic carbide (C4-) at the center of the active site cluster. The role of this carbide is thought to enhance the structural stability, magnetic coupling, and possible stabilization of high oxidation states during catalysis.
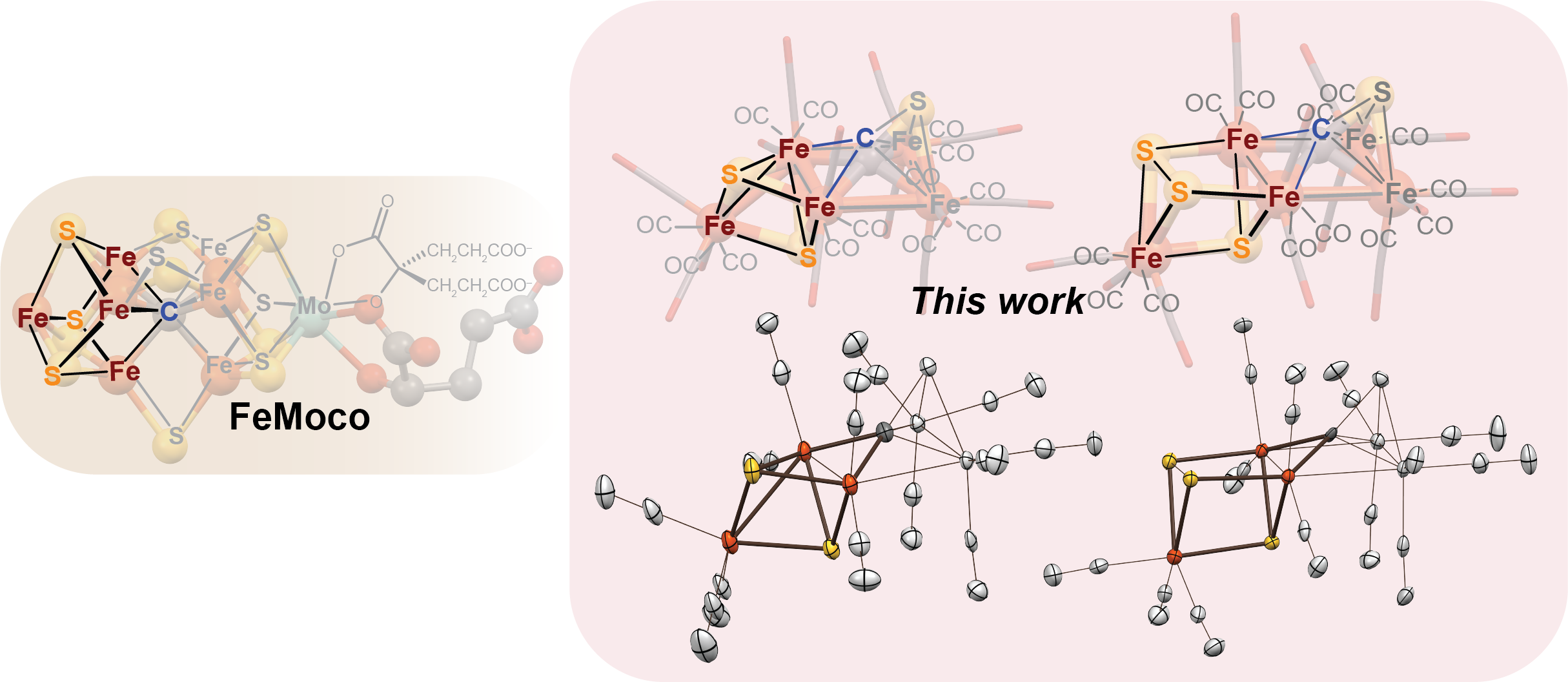
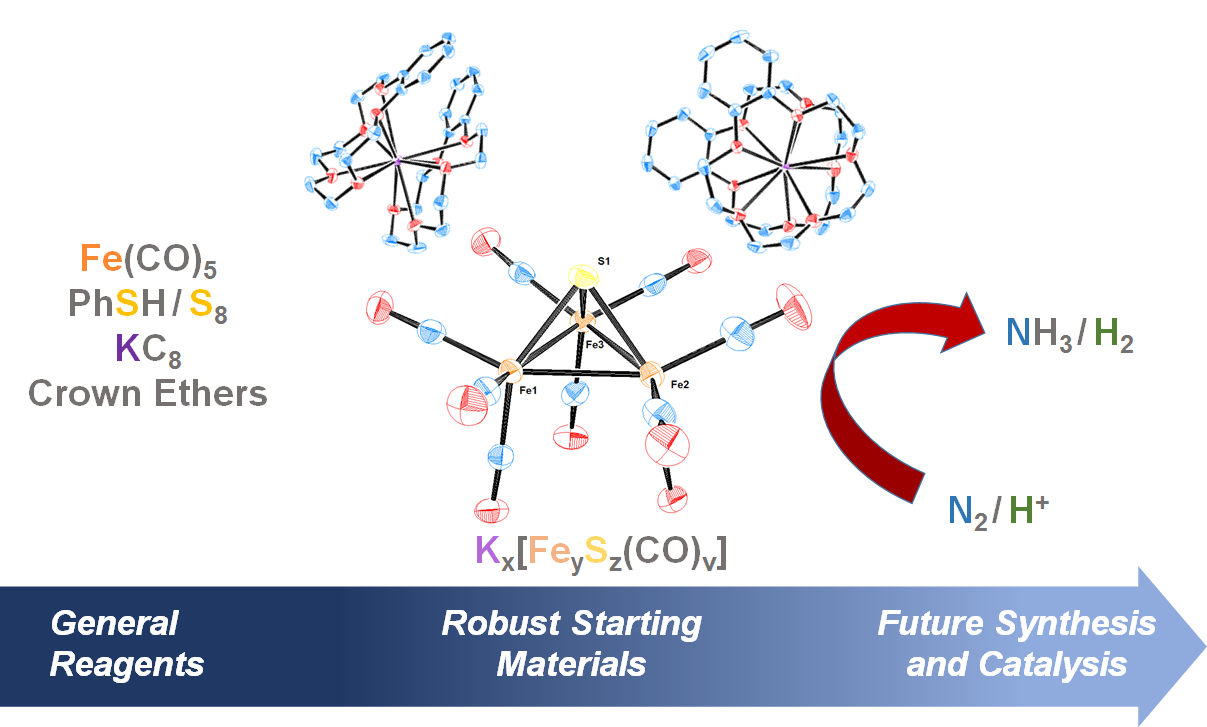
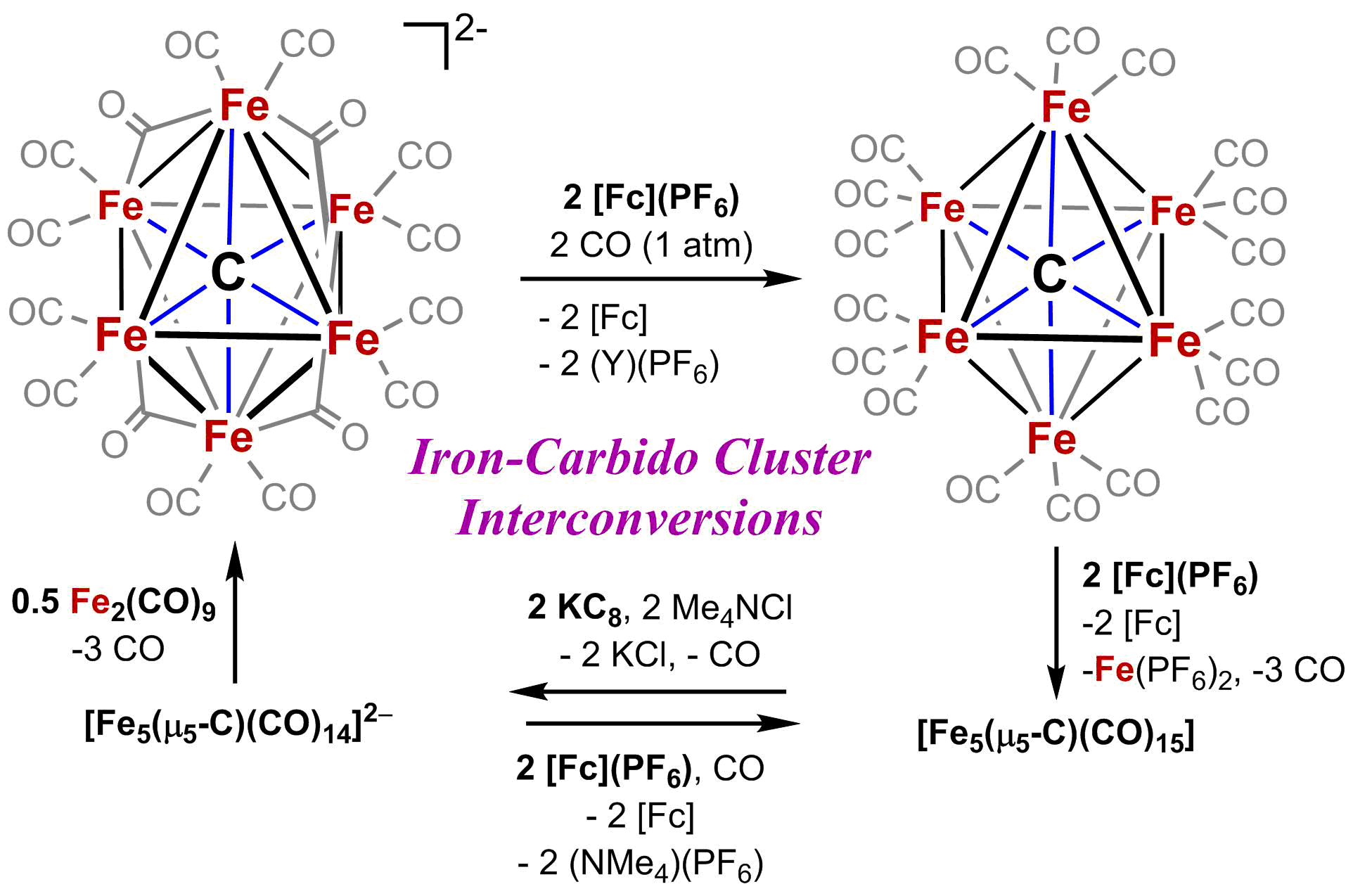
Authentic Iron-Carbide Clusters. However, the bio-inorganic molecular chemistry of the iron-sulfur-carbide cluster is limited by the lack of synthetic models that contain the purely inorganic carbide. We have developed synthetic methods to isolate some of the first Iron-Carbide-Sulfide and the first Iron-Carbide-Thiolate binding motifs (see below, Angew Chem 2021). We are now expanding these methods to develop structurally relevant and catalytically active clusters based on sulfide/phosphide ligation for N2 reduction to ammonia (NH3), as well as unprecedented nitrogen-transfer reactions.
Related Techniques include organic ligand synthesis, inorganic synthesis, X-ray crystallography, and inorganic spectroscopy (EPR, Mossbauer, NMR, UV/vis, XPS) and TD/DFT computations.
Relevant Publications
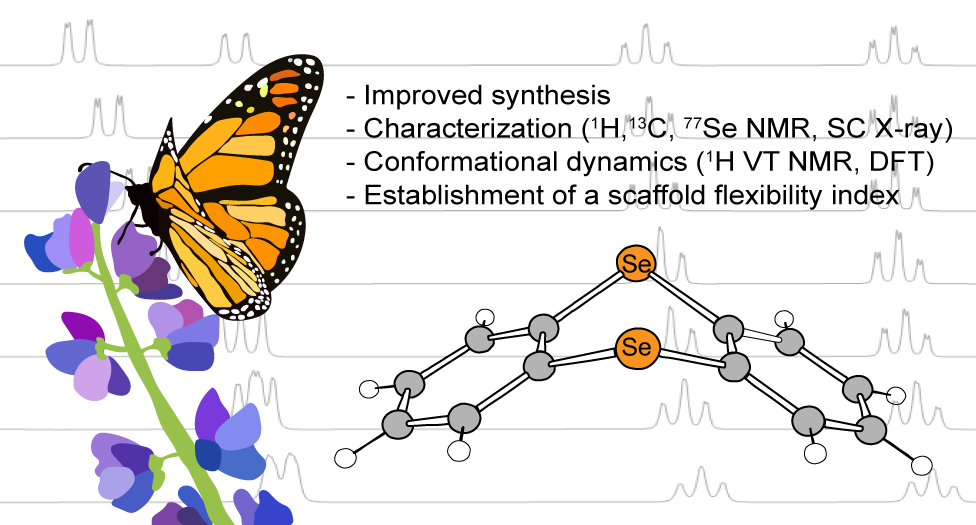
A. T. Larson, B. Boyle, J. Labrecque, A. Ly, C. Bui, S. Vasylevskyi and M. J. Rose. Synthesis & Conformational Dynamics of Selenanthrene (Oxides): Establishing an ‘Energetic Flexibility Index’ for Scaffolds. In Revisions.
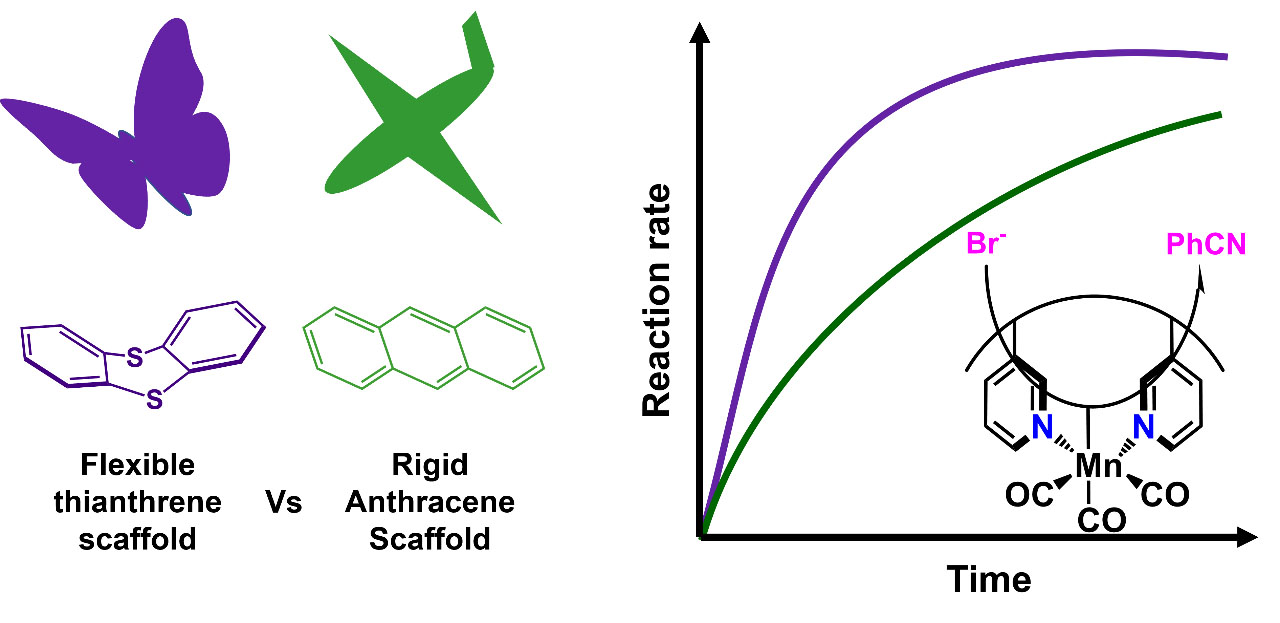
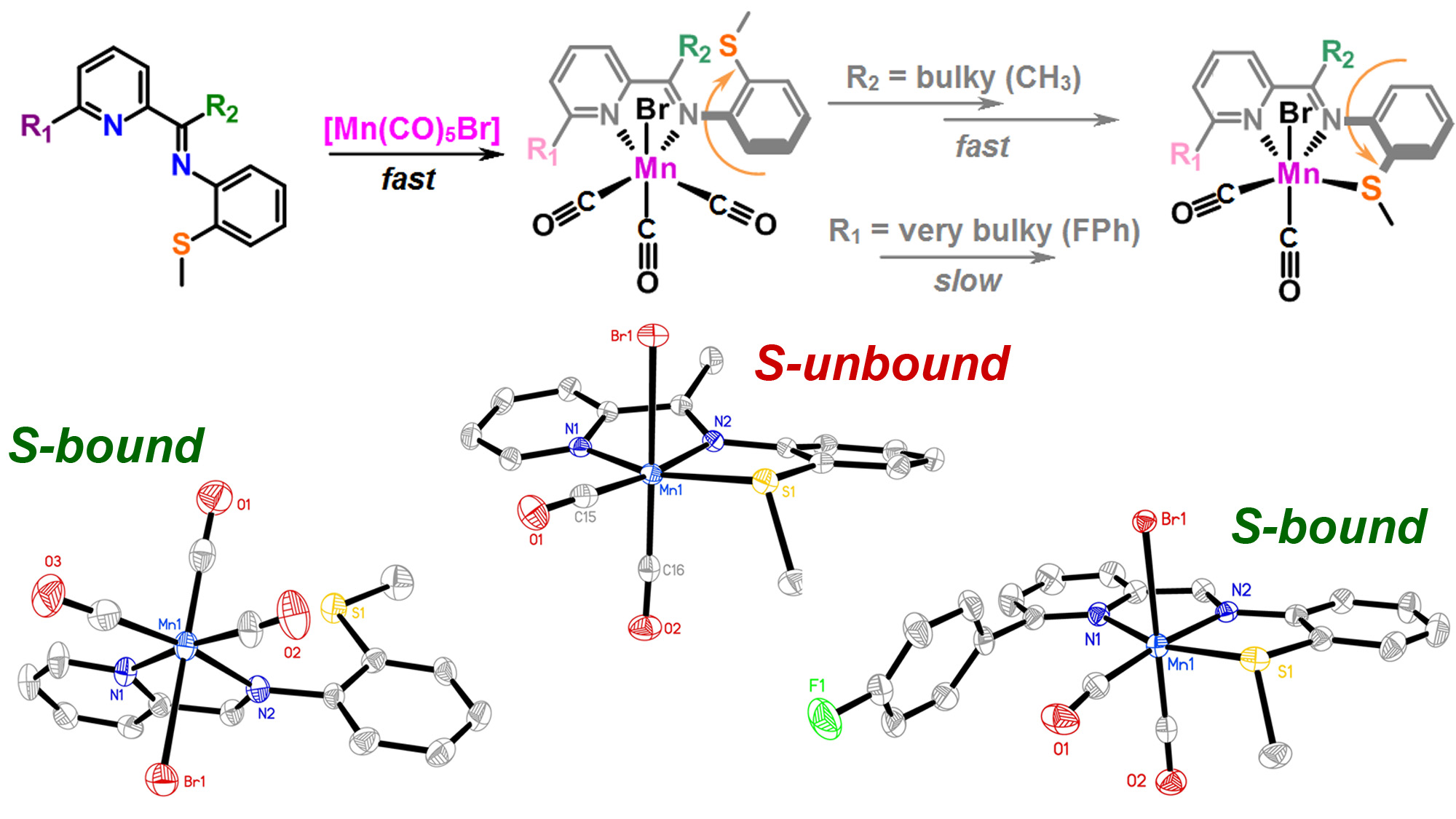
J. Lebrecque, Y. I. Cho, D. K. McIntosh, F. Agboola and M. J. Rose. Distal Scaffold Flexibility Accelerates Ligand Substitution Kinetics in Manganese(I) Tricarbonyls: Flexible Thianthrene versus Rigid Anthracene. Dalton Trans. 2023, 52, 4028-4037. Link
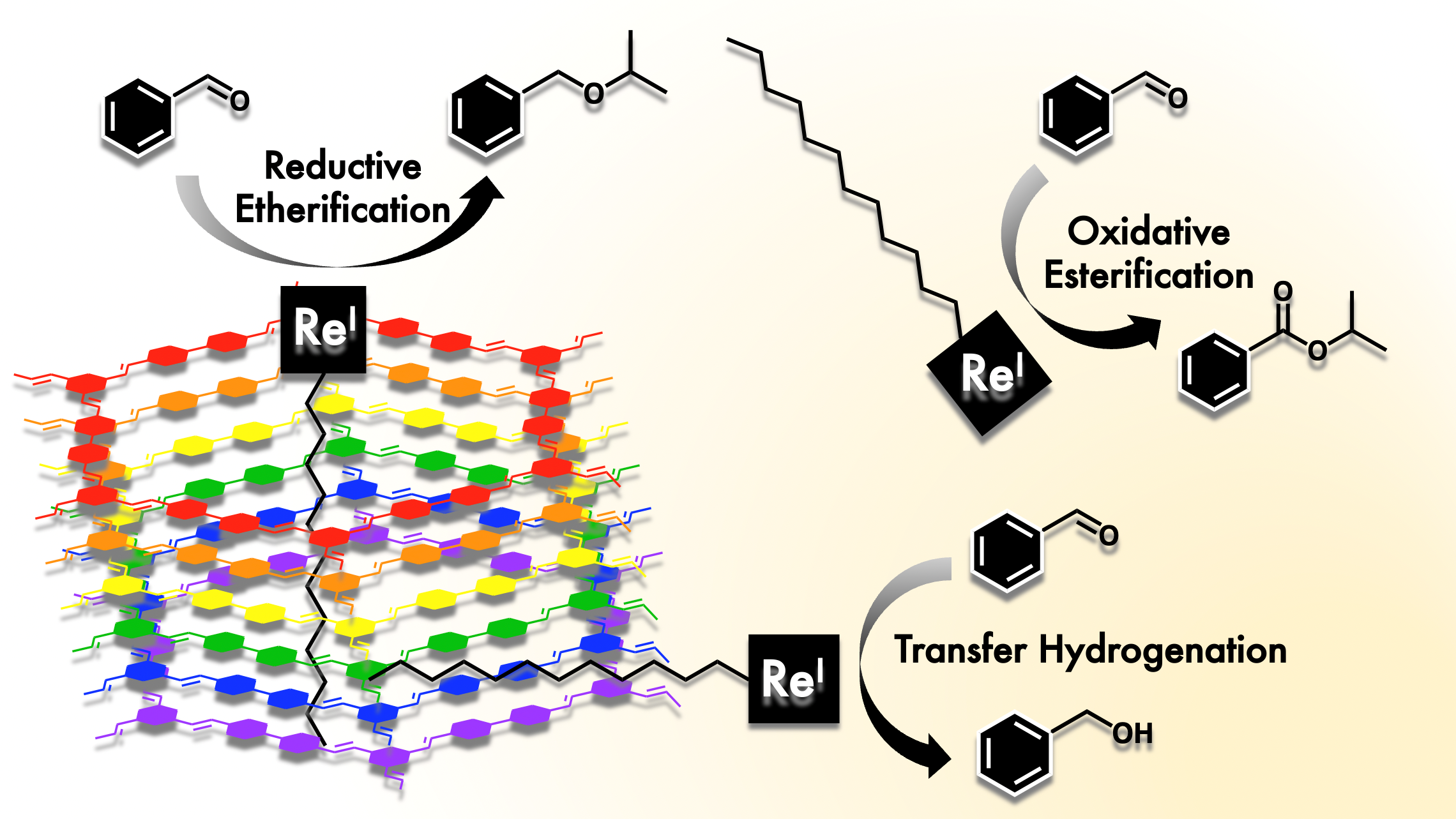
S. T. Goralski, K. M. Cid-Seara, J. J. Jarju, L. Rodriguez-Lorenzo, A. P. LaGrow, M. J. Rose and L. M. Salonen. Threefold Reactivity of a COF-Embedded Rhenium Catalyst: Reductive Etherification, Oxidative Esterification or Transfer Hydrogenation. Chem. Commun. 2022, 12074-12077. Link
S. T. Goralski and M. J. Rose. Emerging Artificial Metalloenzymes Asymmetric Hydrogenation Reactions. Curr. Opin. Chem. Biol. 2021, 102096. Link
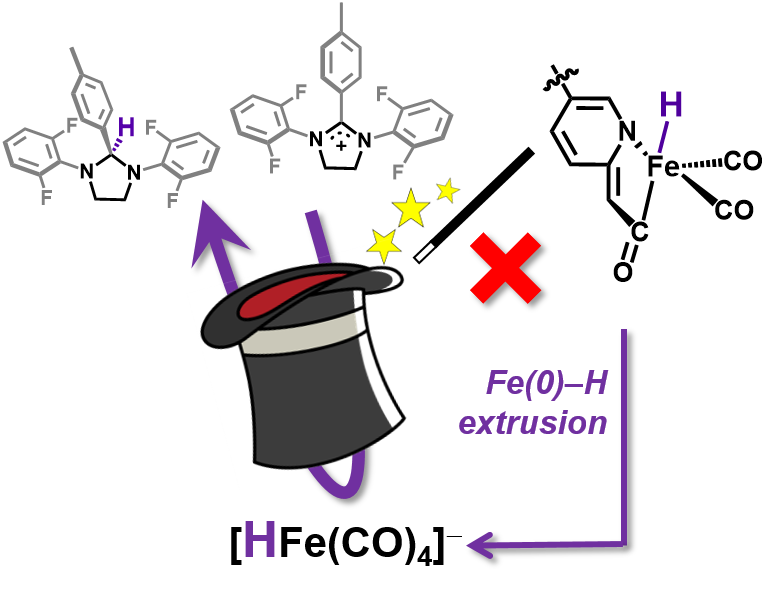
S. A. Kerns#, J. Seo#, V. M. Lynch, J. Shearer, S. T. Goralski, E. R. Sullivan and M. J. Rose. Scaffold-based [Fe]-hydrogenase model: H2 activation initiates Fe(0)-hydride extrusion and non-biomimetic hydride transfer. Chem. Sci. 2021, 12, 12838-12846. Link
C. Joseph, C. R. Cobb and M. J. Rose. Single-Step Insertion of Sulfides and Thiolate into Iron Carbide-Carbonyl Clusters: Unlocking the Synthetic Door to FeMoco Analogues. Angew. Chem. Intl. Ed. 2021, 3475-3479. Link
C. Joseph, J. P. Shupp, C. R. Cobb and M. J. Rose. Construction of Synthetic Models for Nitrogenase Relevant NifB Biogenesis Intermediates and Iron-Carbide-Sulfide Clusters. Catalysts (Bio-Inorganic Special Edition) 2020, 10, 1317. Link
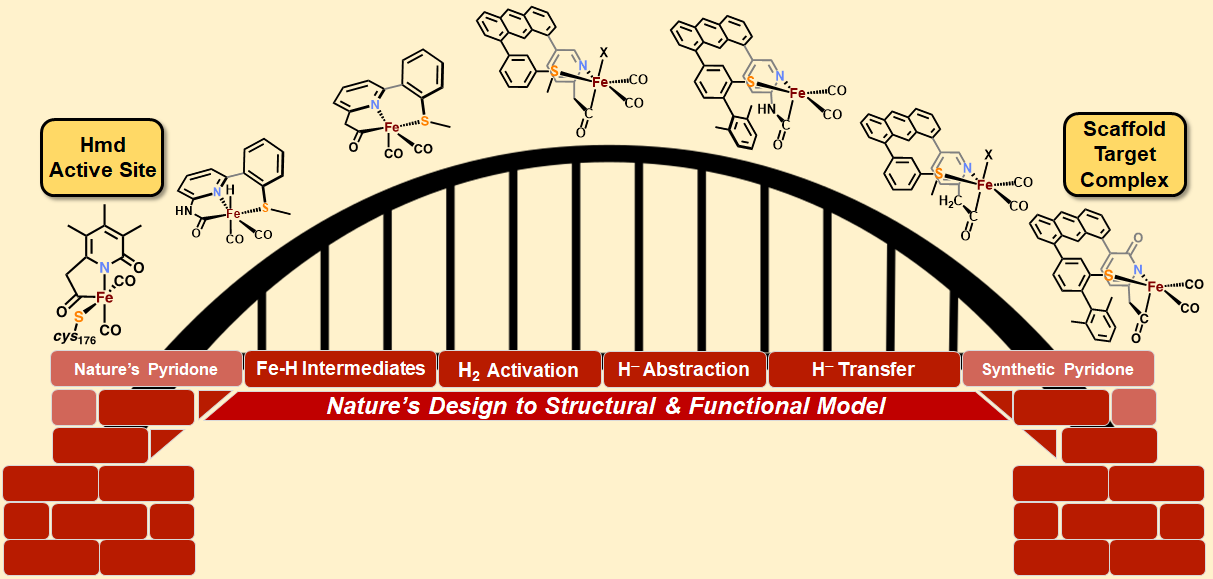
S. A. Kerns and M. J. Rose. Scaffold-based Functional Models of [Fe]-Hydrogenase: Building the Bridge between Biological Structure and Molecular Function. Acc. Chem. Res. 2020, 53, 1637-1647. Link
M. J. Rose. [Fe]-Hydrogenase (Hmd): Insights from Enzyme Structure, Spectroscopy and Synthetic Models. Comprehensive Coordination Chemistry III, 2020. Link
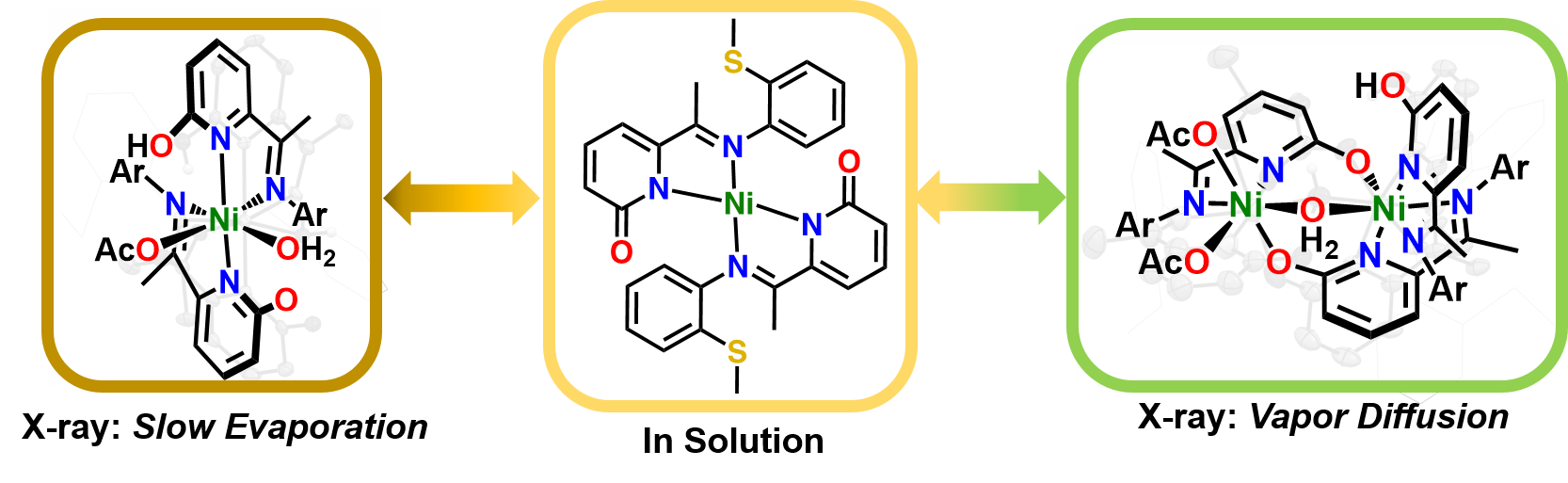
S. T. Goralski, T. A. Manes, S. E. A. Lumsden, V. M. Lynch and M. J. Rose. Divergent Solution and Solid State Structures of Mono- and Dinuclear Nickel(II) Pyridone Complexes. Organometallics 2020, 39, 1070-1079. Link
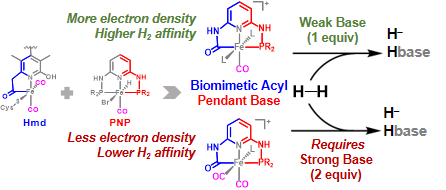
Z.-L. Xie, W. Chai, S. A. Kerns, G. A. Henkelman and M. J. Rose. Bioinspired CNP Iron(II) Pincers Relevant to [Fe]-Hydrogenase (Hmd): Effect of Dicarbonyl versus Monocarbonyl Motifs in H2 Activation and Transfer Hydrogenation. Inorg. Chem. 2020, 59, 2548-2561. Link
J. P. Shupp and M. J. Rose. Facile Hydrogen Atom Abstraction and Sulfide Formation from a Methyl-Thiolate Capped Iron-Sulfur-Carbonyl Cluster. Dalton Trans. 2020, 49, 23-26. Link
J. McGale, G. E. Cutsail III, C. Joseph, M. J. Rose and S. DeBeer. Spectroscopy and Mossbauer Characterization of M6 and M5 Iron(Molybdenum) Carbide Clusters: High Carbide-Iron Covalency Enhances Local Iron Site Electron Density Despite Oxidation. Inorg. Chem. 2019, 58, 12918-12932. Link
Y. I. Cho, G. Durgaprasad and M. J. Rose. CNS and CNP Iron(II) Mono-Iron Hydrogenase (Hmd) Mimics: Role of Deprotonated Methylene(acyl) and the trans-Acyl Site in H2 Heterolysis. Inorg. Chem. 2019, 58, 12689-12699. Link
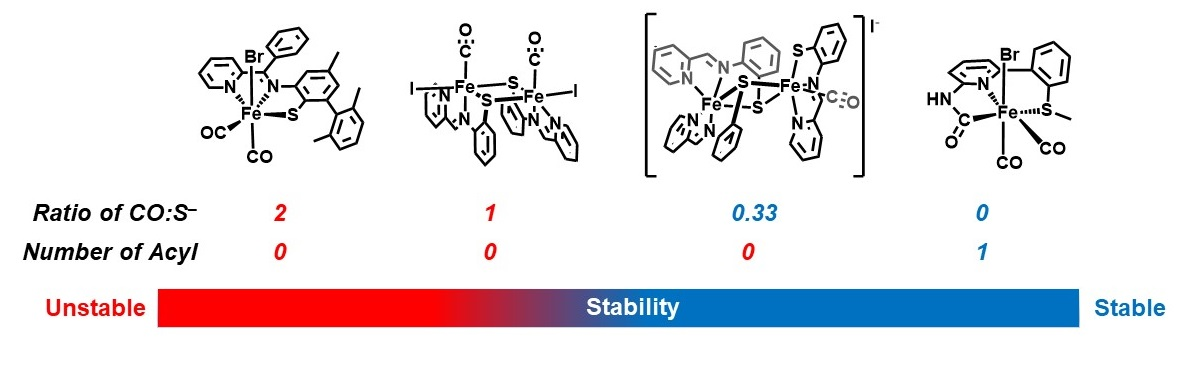
Z.-L. Xie, D. L. Pennington, D. G. Boucher, J. Lo and M. J. Rose. Effects of Thiolate Ligation in Mono-Iron Hydrogenase (Hmd): Stability of the {Fe(CO)2}2+ Core with NNS Ligands. Inorg. Chem. 2018, 57, 10028-10039. Link
C. Joseph, S. Kuppuswamy, V. M. Lynch and M. J. Rose. Fe5Mo Cluster with Iron-Carbide and Molybdenum-Carbide Bonding Motifs: Structure and Selective Alkyne Reductions. Inorg. Chem. 2018, 57, 20-23. Link

S. Kerns, A.-C. Magtaan, P. Vong and M. J. Rose. Functional Hydride Transfer by a Thiolate-containing Model of Mono-iron Hydrogenase featuring an Anthracene Scaffold. Angew. Chem. Int. Ed. 2018, 57, 2855-2858
T. A. Manes and M. J. Rose. Rigid scaffolds for the design of molecular catalysts and biomimetic active sites: A case study of anthracene-based ligands for modeling mono-iron hydrogenase (Hmd). Coord. Chem. Rev. 2017, 353, 295-308. Link
Z.-L. Xie, G. Durgaprasad and M. J. Rose. Substitution Reactions of Iron(II) Carbamoyl-Thioether Complexes Related to Mono-Iron Hydrogenase. Dalton Trans. 2017, 46, 10814. Link
J. P. Shupp, A. R. Rose and M. J. Rose. Synthesis and Interconversions of Reduced, Alkali-Metal Supported Iron-Sulfur-Carbonyl Complexes. Dalton Trans. 2017, 46, 9163-9171. Link
J. Seo, T. E. Sotman, E. R. Sullivan, B. D. Ellis, T. Phung and M. J. Rose. Structural and Electronic Modifications of Pyridones and Pyrones via Regioselective Bromination and Trifluoromethylation. Tetrahedron 2017, 73, 4519-4528. Link

S. Kuppuswamy, J. Wofford, C. Joseph, A. K. Ali, V. M. Lynch, P. A. Lindahl, and M. J. Rose. Structures, Interconversions and Spectroscopy of Carbonyl Clusters with an Interstitial Carbide: Localized Iron Center Reduction via Cluster Oxidation. Inorg. Chem. 2017, 56, 5998-6012. Link
J. Seo, T. A. Manes and M. J. Rose. Structural and Functional Synthetic Model of Mono-Iron Hydrogenase Featuring an Anthracene Scaffold. Nature Chem. 2017, 9, 552-557. Link
T. A. Manes and M. J. Rose. Mono- and Di-nuclear Manganese Carbonyls Supported by 1,8-Disubstituted (L = Py, Ar-SMe, Ar-SH) Anthracene Ligand Scaffolds. Inorg. Chem. 2016, 55, 5127-5138. Link
G. Durgaprasad, Z.-L. Xie and M. J. Rose. Iron Hydride Detection and Intramolecular Hydride Transfer in a Synthetic Model of Mono-Iron Hydrogenase with a CNS Chelate. Inorg. Chem. 2016, 55, 386-389. Link
Y. I. Cho, M. L. Ward and M. J. Rose*. Substituent Effects of N4 Schiff Base Ligands on the Formation of Fluoride-Bridged Dicobalt(II) Complexes via B–F Abstraction: Structures and Magnetism. Dalton Trans. 2016, 45, 13466-13476.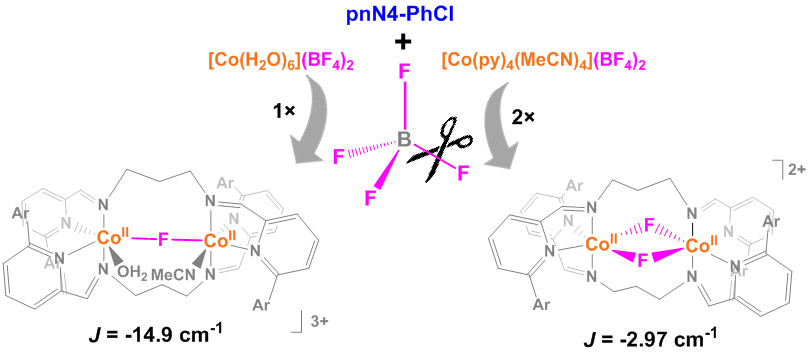
K. A. Thomas Muthiah, G. Durgaprasad, Z.-L. Xie, O. M. Williams, C. Joseph, V. M. Lynch and M. J. Rose. Mononuclear Iron(II) Dicarbonyls Derived from NNS Ligands: Structural Models Related to a Possible “Pre-Acyl” Active Site of Mono-Iron (Hmd) Hydrogenase. Eur. J. Inorg. Chem. 2015. 1675-1691. Link
J. Seo, A. Ali, M. J. Rose. Novel Ligand Architectures for Metalloenzyme Modeling: Anthracene-based Ligands for Synthetic Modeling of Mono-[Fe] Hydrogenase. Comments Inorg. Chem. 2014, 34, 103-113. Link
S. E. A. Lumsden, G. Durgaprasad, K. A. Thomas Mutiah, M. J. Rose. Tuning Coordination Modes of Pyridine/Thioether Schiff Base (NNS) Ligands to Mononuclear Manganese Carbonyls. Dalton Trans. 2014, 43, 10725. Link
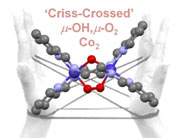
Y. I. Cho, D. M. Joseph and M. J. Rose. ‘Criss-Crossed’ Dinucleating Behavior of an N4 Schiff Base Ligand: Formation of a mu-O2, mu-OH Dicobalt(III) Core via O2 Activation. Inorg. Chem. 2013, 52, 13298-13300. Link